Table of Contents
Data source
The data used in this study are from the Danish national health registers. All health data is reported when a patient among others is hospitalized, visits a GP, or picks up medication at the pharmacy. Other registers contain information on demographics and socioeconomic status. All data is stored safely and is not accessible to the public. Each citizen in Denmark is identifiable by a Social Security number (CPR). This CPR number is encrypted to a unique personal identification number (PNR). PNR is a key, that enables linkage of data across registers. Table 1 illustrates the specific database.
The outcomes, nephropathy, tissue infection and cardiovascular events are defined by ICD-10 codes from The National Patient Register (LPR). The specific ICD-10 codes used can be found in supplementary Table S3.
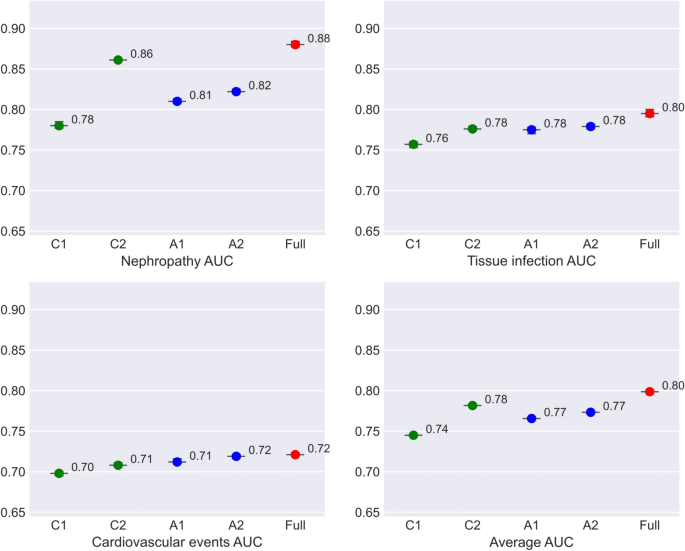
For each outcome, we develop five different models in stages to evaluate the model performance, using different data sources. The specifications of the five models are specified in Table 2.
Having comorbidities and multimorbidity are well known risk factors for disease progression. The Charlson Comorbidity Index (CCI) is based on ICD10 diagnoses19 and is one of the most used measures for morbidity20. We include CCI as a binary indicator equal to 1 if a patient have had one of the included ICD10 diagnoses in the past five years. To improve coverage of morbidity signals for the prediction we add features from alternative measure of morbidity by also including features based on the Elixhauser Comorbidity Index (ECI)21. This index is also based on ICD10 diagnoses and we add binary features for presence of congestive heart failure, historic registration of depression, and registered obesity
Cohort and exclusion criteria
In Denmark, T2D is defined by The Danish Health Data Authority, who have developed an algorithm for selected chronic diseases and severe mental disorders called RUKS22. Using the RUKS algorithm23, all PNR on T2D patients are obtained in the period 2014-2019. This makes it possible to link all health information on these patients through PNR, in the different registers.
Laboratory test results were only accessible for regions of Southern Denmark, Northern Jutland and Zealand in the defined time period. This means that patients with diabetes living in the capital region and the region of Central Jutland are excluded, effectively cutting the patient population in half. Using this cohort, we end up with 104,341 eligible unique patients with diabetes. Descriptive statistics of the cohort characteristics are in the supplementary materials, Table S4, as well as the descriptive statistics of training, validation, and test sets.
Study design
The dataset consists of instances, each instance is an observation period, a buffer period and a target window. This has been applied for each eligible patient, such that multiple observations for a given patient are possible. Each instance is offset by three months, so that target windows are non-overlapping. This approach is illustrated in Figs. 4 and 5, and has been used in similar prior studies6,24.
Information Extraction Shows the structure of a single instance, and how the information is extracted. The observation period is one year, here the information is extracted. The buffer is two years. And finally, the target window, where the complication is predicted.

Data Point creation Shows how each data point/instance is created. Each blue and orange represents a sample in the data and is explained in Fig. 4. For instance, Patient 1 might have 5 different instances. Each instance is added to the dataset as a different data point. Patient 2 only has one instance, this could be due to death post first data point, therefore additional data points are not created. For patient 3, there is two data points. The late entry could be due to later diagnosis.
For the observation period, we used one year of data, which is sufficient to capture the relevant characteristics of each instance. This choice is further supported by clinical practice, where an HbA1c test is recommended at least once a year25.
The buffer period is the period between observation and prediction. The buffer period is set to two years, meaning that the prediction is two years post the observation window. This makes it possible to make a prediction of the two-year risk of adverse outcomes. The two-year buffer is chosen since a possible adverse outcome of complication in some cases is avoidable if intervened or decelerated, depending on the complication.
The target window of outcome prediction, ’complication prediction’, is 3 months following the buffer. With this data structure, a patient can generate up to 8 instances. In the case where the data does not allow a complete instance, the instance is excluded.
An example of the first two instances of a patient could be: instance 1: Observation period from January 2014 to December 2014 a buffer from January 2015 to December 2016 and the target window from January 2017 to March 2017. Instance 2: Observation period April 2014 to March 2015, a buffer from April 2015 to March 2017, and the target window from April 2017 to June 2017.
The data is randomly split into three different datasets on the individual level. All instances belonging to an individual are in the same dataset. For instance the Fig. 5, all instances of patient 1 are in training data, all instances of patient 2 are in validation, and all instances of patient 3 are in the test.
Model development
The project was implemented in Python version 3.10.10. Key libraries utilized include xgboost (v1.7.5) for model training, shap (v0.41.0) for explainability analysis, and dalex (v1.6.0) for model fairness assessment. Other core packages can be found in supplementary Table S5.
We use XGBoost, an efficient and powerful machine learning algorithm, for our predictions26. XGBoost is widely applied in studies utilizing tabular data, demonstrating robust performance on tabular applications6,24. For tasks such as this, it has been proven to be a well-performing algorithm, even when compared to deep learning techniques27. Additionally, it is able to handle missing values, and as laboratory test results tend to have a rather large frequency of missing values – as some patients do not get tested – this is necessary. Missing values might even provide useful information28, and could therefore recognize these behavioral patterns.
Since the distribution of classes is imbalanced for all outcomes due to the low frequency of diabetes complication in any 3-month period, it is necessary to balance the data to avoid biased predictions and poor performance. To do so, the training data are undersampled such that the dimension of negative cases are equivalent to the positive cases. Undersampling was applied exclusively to the training data. The validation and test datasets were preserved with their original class distribution.
To evaluate the performance of the trained models, each model was retrained five times using random restarts. We do this to account for differences in the undersampled training set introduced by the random undersampling of negative cases. By repeating the training process multiple times, we aim to address the differences and report a more robust model performance.
To find the most optimal hyperparameters in the XGBoost model29, we used a grid search approach to find the parameters that resulted in the most optimal model. For each model and each outcome, the most optimal hyperparameters that maximized the validation AUC were found using the python module Hyperopt30. The final hyperparameters for each model and outcome are presented in supplementary Table S6.
Feature importance analysis
To investigate which features contribute most to the model predictions, we analyzed feature importance across the different outcomes. Feature importance was calculated using SHAP (SHapley Additive exPlanations) values in python module shap. We computed the mean absolute SHAP values for each feature, averaged over five independent runs to ensure stability and robustness. To estimate the feature contributions, we used the test data. The top six features, ranked by their average SHAP values, are reported for each model and outcome in Fig. 2. Additional SHAP analyses are in the supplementary materials, Fig. S2.
Evaluating the algorithmic fairness of the models
To investigate potential disparities in machine learning models, we compare various performance metrics using the confusion matrix, supplementary Fig. S3: True Positive Rate (TPR) \(\frac{TP}{TP+FN}\)False Positive Rate (FPR) \(\frac{FP}{FP+TN}\)Predictive parity ratio (PPV) \(\frac{TP}{TP+FP}\)Statistical Parity (STP) \(\frac{TP+FP}{TP+FP+TN+FN}\)and Accuracy (ACC) \(\frac{TP+TN}{TP+FP+TN+FN}\). We analyze these metrics for one subgroup (e.g., males) in comparison to its counterpart (e.g., females) to gain insights into model performance across these subpopulations.
A feature is considered biased if the ratio of performance metrics between two groups falls outside a specified range. When comparing males and females, we determine a model to be fair if this ratio lies within the range: \(\epsilon< \frac{metric_{female}}{metric_{male}} < \frac{1}{\epsilon }\). Where \ (\ Epsilon \) is set to 0.8, as this is the most commonly used threshold, and corresponds to the four-fifths rule31.
A machine learning model may exhibit discrimination against a subgroup if there is encoded bias within the data, if there is missing or imbalanced data, or if the model relies heavily on sensitive attributes. Biases in the data may stem from historical inequalities, non-representative sampling, or societal stereotypes, which can subsequently be learned and perpetuated by the model32,33. For prediction using health data, it is also possible that two subgroups are biologically different and therefore experience different prognoses.